Research Strategy
Most of the basic ideas of ‘systems biology’ remain to be applied to relevant biological problems.
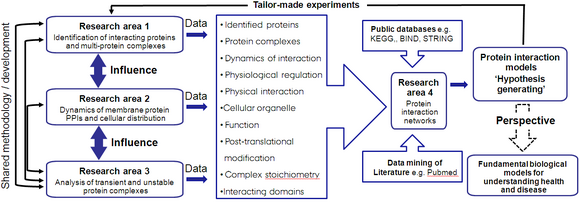
InterPrET focuses on obtaining detailed understanding of how PPIs regulate specific types of membrane proteins, each with profound biological and medical relevance (see Research areas). The research programs within InterPrET are divided into research areas (RAs), each designed to address individual center objectives. Although the RAs operate independently to provide essential biological data; they are not fixed entities. Each RA is highly interlinked and flexible; e.g. data from one RA can directly influence the direction of other RAs, there is an unrestricted flow of ideas and shared method/technique development (see figure):
- RA1: Identification of interacting proteins and multi-protein complexes. The major aim in this area is the development of state-of-the art techniques for the identification of intact membrane protein complexes.
- RA2: Functional protein:protein interactions. The focus within this area is the development and use of an array of advanced live cell imaging techniques to determine the dynamics and location of PPIs and protein complex formation.
- RA3: Analysis of transient protein complexes. Unstable, macromolecular membrane protein assemblies have so-far escaped structural characterization due to a lack of suitable methods; we are currently developing novel approaches to analyze instable membrane protein complexes.
- RA4: Modeling of protein interaction networks. To put results into context, bioinformatics/modeling studies are underway to uncover essential protein networks for specific cellular processes.
How is the InterPrET approach different? In eukaryotes, the majority of previous studies for analysis of protein interactions and protein networks have been performed in ‘model organisms’. These approaches have provided pieces to the puzzle, but they have not put them together to produce a coherent picture that is physiologically relevant. For example, some large-scale approaches used do not measure interactions between proteins in their natural cellular context and are not amenable to studying protein complexes that are dynamic, transiently associated under different conditions and regulated by a physiological stimulus.
InterPrETs approach will overcome these problems by focusing on PPI-mediated regulation of a small set of biologically and medically important membrane proteins from multiple angles. By joining researchers with complementary but diverse techniques we can comprehensively evaluate regulated membrane PPIs at multiple levels; from complex identification, sorting and localization, through to function. This integrated approach will provide all the necessary data in one system to model biologically relevant PPIs and build fundamental regulatory mechanisms for other proteins and/or cell types.